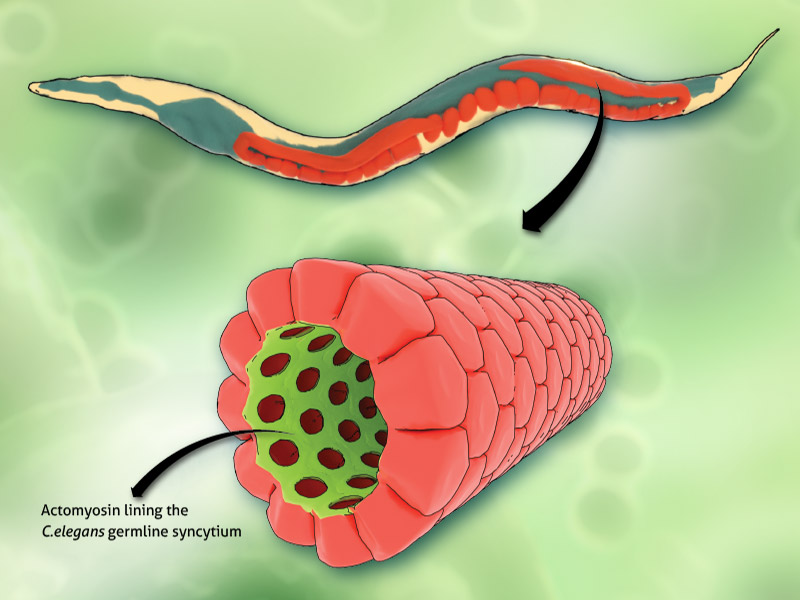
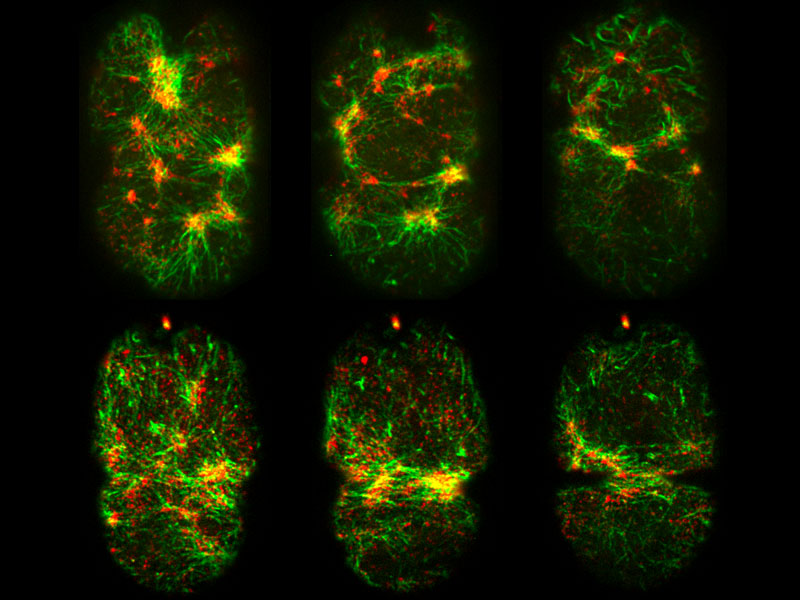
Microscopic Muscles
Steven j wolf, phd | march 2017
Researchers from the Mechanobiology Institute, National University of Singapore have described, for the first time, the ordered arrangement of myosin-II filaments in actin cables of non-muscle cells. This work is published in Nature Cell Biology (Hu S et al. Long-range self-organization of cytoskeletal myosin II filament stacks. Nature Cell Biology, 19, 133–141 (2017) doi: 10.1038/ncb3466).
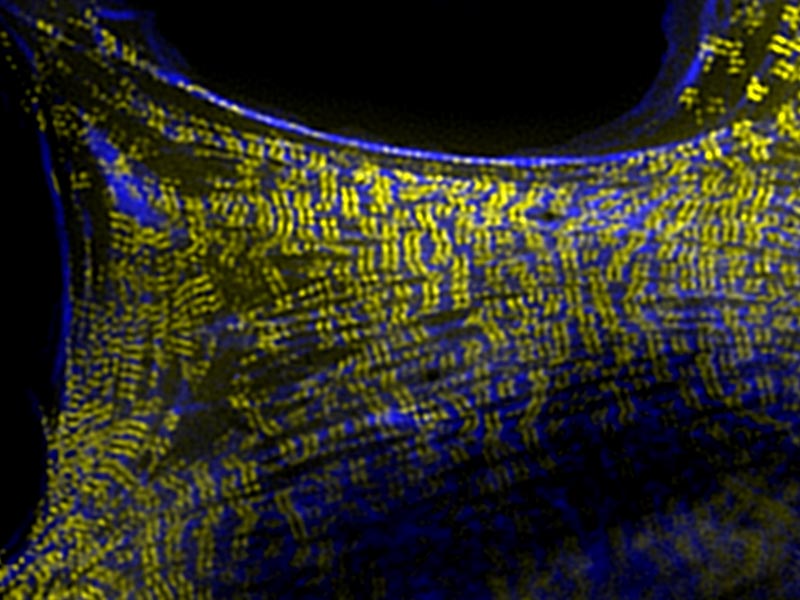
Visualization of myosin II filament stacks (yellow) with cross-linking protein alpha-actinin-1 (blue) in a rat embryo broblast (REF52) cells using structured illumination microscopy (SIM, Nikon). Myosin II filaments alternate with alpha-actinin enriched domains along actin stress fiber.
How non-muscle cells find the strength to move
The twitching contractions of our muscle cells are well known. They can be detected just weeks after conception as the embryonic heart begins beating. Muscle cell contractility is produced from interactions between protein-based cables of the cytoskeleton and small molecular motor proteins known as myosins.
There are over 200 types of cells within the human body, and not all need to repeatedly contract. Despite their distinct functions, nearly all cells contain the same basic protein components found in muscle cells. Importantly, most cells also exhibit some degree of slow contractility. Fibroblasts are one such example. Found in connective tissue, these cells produce the material that surrounds all cells, and ultimately defines tissue shape. Importantly, fibroblasts are also known to remodel this material, and for this, they need strength to pull against their environment.
To investigate the organization of the cytoskeleton and its associated motor proteins in non-muscle cells, researchers from the Mechanobiology Institute, National University of Singapore, analyzed fibroblasts using a form of super resolution microscopy known as Structured Illumination Microscopy (SIM).
The researchers, who were led by Prof. Alexander Bershadsky and Asst. Prof. Ronen Zaidel-Bar, focused their investigation on the assembly of the cytoskeleton. Along with providing structural support to the cell, the cytoskeleton can also buffer stresses from the external microenvironment and give cells the power to contract and move through a tissue. These processes are possible due to the continual assembly and disassembly of the protein cables, and due to the generation of force as motor proteins pull on these cables.
When the cytoskeleton was viewed in living fibroblasts, Dr Shiqiong Hu, a postdoctoral researcher at MBI, and colleagues, discovered unique, organized patterns of motor protein filaments within large protein cable-like structures known as stress-fibers. These cables form dynamically and often bridge sites where the cells are interacting with the microenvironment.
Ordered arrangement of myosin-II filaments defined in non-muscle cells
Like ropes, these cables are made up of many individual filaments, held together by various cross-linking proteins. By watching the cytoskeleton form over time, the researchers observed how myosin-II filaments arranged into stacks that ran perpendicular to the large parallel stress fibers. These stacks alternated with regions of the ‘cross-linking’ protein a-actinin, which tethers individual filaments together to produce the protein cable.
How myosin-II filaments come to be stacked together within the bundled stress fibers, remains to be fully defined, however one observation from this study that may hold the answer, is the long range movement of myosin-II filaments towards each other. As the researchers propose, this attraction may result from contractile or elastic forces generated by the myosin filament stacks, which can transmit through the surrounding cytosol to individual filaments that are otherwise isolated.
The stacking of myosin-II filaments in non-muscle cells like the fibroblast is an intriguing element in the self-organization of the cytoskeleton, and overall architecture of the cell. Fibroblast function requires the cell to be able to stretch, generate cytoskeletal protrusions and move to other regions of the connective tissue. The assembly and organization of myosin-II into stacks permits the fibroblast to fulfill these cellular processes.
Even in non-muscle cells the architecture of the cytoskeleton is specialized for force generation and sensing. The organization of the cytoskeleton in non-muscle cells is strikingly similar to that in muscle cells. In both cases contractile and elastic forces are integral in establishing a functional cytoskeleton, and once formed, a pattern of repeating protein-based contractile proteins becomes evident. However, unlike in muscle cells, these structures continuously assemble and disassemble in non-muscle cells, allowing them to adapt their function, shape, and direction of movement according to the environment they find themselves in.
As observed in this study, even non-muscle cells require the strength to pull against their surroundings, and fight their way through often sticky environments. This strength comes from a highly refined system of filaments and motor proteins. Although not as strong as those found in muscle cells, their organization in non-muscle cells allows them to remain responsive to changes in the environment, whilst providing just the right amount of force to carry out their functions.