What is the role of formin in actin polymerization?
Formins promote the elongation of pre-existing filaments by removing barbed end capping proteins and forming a sleeve around the actin subunits. Formins are also capable of actin nucleation, a process which is spatiotemporally coupled with actin disassembly [1].
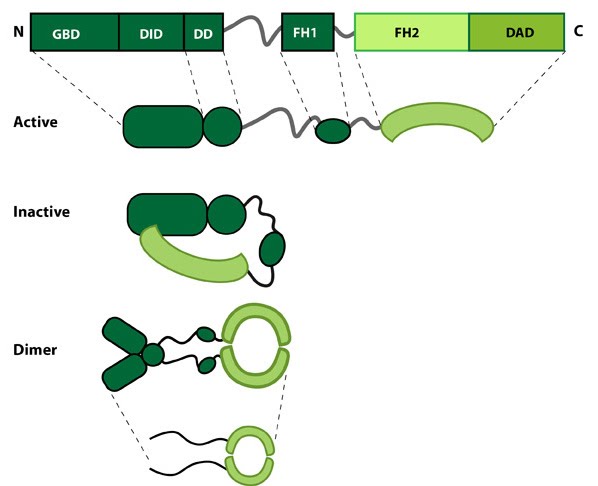
This schematic diagram illustrates the molecular organization of formin. Relevant domains/regions that are believed to be important for actin binding and protein-protein interactions are highlighted (reviewed in [2]. An intramolecular interaction between the diaphanous inhibitory domain (DID)and the diaphanous auto-regulatory domain (DAD), which prevents formin (e.g. mDia1) from nucleating actin filaments, is relieved by Rho binding to the GTPase binding domain (GBD, aka CRIB domain) [3]. However, this regulation may be more complex [4, 5]. DD = dimerization domain, FH = formin homology domain
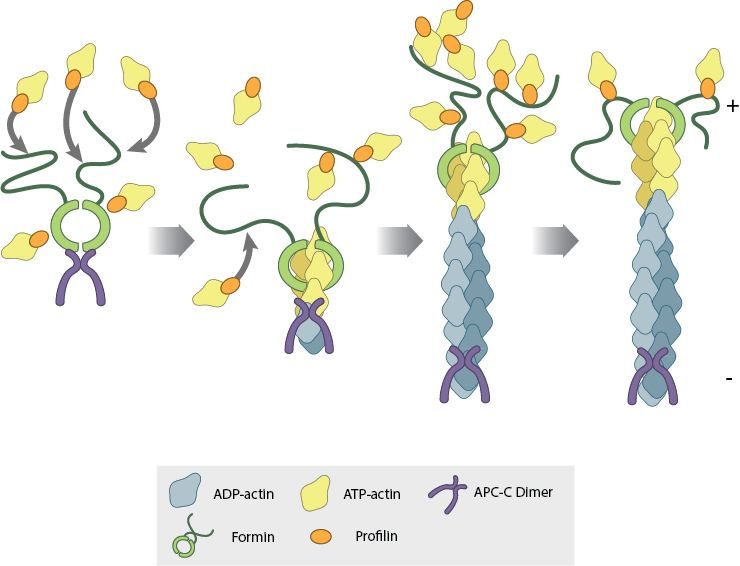
The FH2 domains of the formin dimer (shown in green) bind to actin monomers to initiate filament assembly. Recent studies indicate this is assisted, or even mediated, by additional factors such as APC. The FH1 domains of the formin dimer (shown as black lines) have short polyproline sequences that interact with profilin. Profilin binds to both formin and actin monomers to increase the addition of actin monomers to the barbed end of the filament.
It is well-established that activated formins facilitate elongation as dimers and form a donut-shaped complex around terminal actin subunits, orientating themselves toward the (+) end of the actin filament [2]. Binding, which is facilitated by FH2 (formin homology 2) domains within the formin monomers removes capping protein from the end of the filament and prevents re-capping to allow continued growth of filaments or cross-linked bundles ([3], reviewed in [4]).
A role in actin filament nucleation
Formins nucleate and polymerize actin filaments at focal adhesions at a rate of around 0.3 µm/min [5]. Inhibiting formin protein expression results in a decreased filament elongation rate (0.1 µm/min), coupled with abnormal stress fiber morphology and an accumulation of actin binding proteins (e.g. α-actinin [5]). Ena/VASP proteins support formin-mediated filament elongation by tethering the filaments near sites of active actin assembly [6][7].
Next, each formin monomer binds and captures profilin units, which are themselves already bound to G-actin monomers. This interaction is mediated by multiple stretches of polyproline residues within the FH1 domain of formins [8]. This domain is known to range from 15-229 residues, consist of between 35% and 100% proline residues, and contain up to 16 profilin binding sites [9]. Profilin maintains a steady pool of actin monomers by promoting ADP to ATP nucleotide exchange on G-actin [10]. These monomers of ATP-G-actin are then added the growing actin filament. The coupling of formin with the growing end prevents capping and allows continued growth of the filaments [11].
Formins could, in theory, contribute to protrusive forces by remaining attached to the barbed end of actin filaments [12][13] (reviewed in [9]). Consistent with this notion, excessive formin activity promotes cell migration. However, the specific mechanisms involved remain unknown given that formin-induced activity does not impact the overall adherence of cells to their substrate, nor does it change the avidity or affinity of cell adhesion receptors (e.g. integrins) [14].
What is the role of profilin in formin-nucleated actin cable assembly?
Profilin binds simultaneously to formin and actin monomers; this interaction tethers multiple profilin-actin complexes near the growing end of actin filaments, which promotes the processive addition of actin subunits [15][16]. Profilin uses the energy from ATP hydrolysis generated during actin polymerization to facilitate actin assembly [16]. Profilin binds to cytoplasmic ATP-actin monomers better than cytoplasmic ADP-actin monomers [17].
Profilin has been suggested to generally increase the elongation rate of formin-associated filaments by:
- Catalyzing the exchange of ADP for ATP on actin monomers [18][19].
- Blocking free monomers from elongating pointed ends [18].
- Lowering the critical concentration at the barbed end [16].
- Promoting the association of G-actin-ATP to the barbed end [20].
Importantly however, profilin also promotes disassembly of actin filaments by sequestering monomeric G-actin, thereby blocking its association with the barbed ends and promoting its disassembly from the pointed ends of actin filaments [18]. The combined actions of profilin and ADF/cofilin synergize to enhance turnover of actin filaments [21].
References
- Watanabe N. Inside view of cell locomotion through single-molecule: fast F-/G-actin cycle and G-actin regulation of polymer restoration. Proc. Jpn. Acad., Ser. B, Phys. Biol. Sci. 2010; 86(1):62-83. [PMID: 20075609]
- Markov IN. [Diagnosis of pancreatic insuloma]. Khirurgiia (Mosk) 1978;(11):79-86. [PMID: 214622]
- Yang C, Czech L, Gerboth S, Kojima S, Scita G, and Svitkina T. Novel roles of formin mDia2 in lamellipodia and filopodia formation in motile cells. PLoS Biol. 2007; 5(11):e317. [PMID: 18044991]
- Chesarone MA, DuPage AG, and Goode BL. Unleashing formins to remodel the actin and microtubule cytoskeletons. Nat. Rev. Mol. Cell Biol. 2009; 11(1):62-74. [PMID: 19997130]
- Hotulainen P, and Lappalainen P. Stress fibers are generated by two distinct actin assembly mechanisms in motile cells. J. Cell Biol. 2006; 173(3):383-94. [PMID: 16651381]
- Schirenbeck A, Arasada R, Bretschneider T, Schleicher M, and Faix J. Formins and VASPs may co-operate in the formation of filopodia. Biochem. Soc. Trans. 2005; 33(Pt 6):1256-9. [PMID: 16246092]
- Breitsprecher D, Kiesewetter AK, Linkner J, Urbanke C, Resch GP, Small JV, and Faix J. Clustering of VASP actively drives processive, WH2 domain-mediated actin filament elongation. EMBO J. 2008; 27(22):2943-54. [PMID: 18923426]
- Paul AS, Paul A, Pollard TD, and Pollard T. The role of the FH1 domain and profilin in formin-mediated actin-filament elongation and nucleation. Curr. Biol. 2007; 18(1):9-19. [PMID: 18160294]
- Higgs HN. Formin proteins: a domain-based approach. Trends Biochem. Sci. 2005; 30(6):342-53. [PMID: 15950879]
- Romero S, Le Clainche C, Didry D, Egile C, Pantaloni D, and Carlier M. Formin is a processive motor that requires profilin to accelerate actin assembly and associated ATP hydrolysis. Cell 2004; 119(3):419-29. [PMID: 15507212]
- Pring M, Evangelista M, Boone C, Yang C, and Zigmond SH. Mechanism of formin-induced nucleation of actin filaments. Biochemistry 2003; 42(2):486-96. [PMID: 12525176]
- Firat-Karalar EN, and Welch MD. New mechanisms and functions of actin nucleation. Curr. Opin. Cell Biol. 2010; 23(1):4-13. [PMID: 21093244]
- Dominguez R. Structural insights into de novo actin polymerization. Curr. Opin. Struct. Biol. 2010; 20(2):217-25. [PMID: 20096561]
- Mizuno H, Higashida C, Yuan Y, Ishizaki T, Narumiya S, and Watanabe N. Rotational movement of the formin mDia1 along the double helical strand of an actin filament. Science 2010; 331(6013):80-3. [PMID: 21148346]
- Watanabe N, Madaule P, Reid T, Ishizaki T, Watanabe G, Kakizuka A, Saito Y, Nakao K, Jockusch BM, and Narumiya S. p140mDia, a mammalian homolog of Drosophila diaphanous, is a target protein for Rho small GTPase and is a ligand for profilin. EMBO J. 1997; 16(11):3044-56. [PMID: 9214622]
- Pantaloni D, and Carlier MF. How profilin promotes actin filament assembly in the presence of thymosin beta 4. Cell 1993; 75(5):1007-14. [PMID: 8252614]
- Perelroizen I, Didry D, Christensen H, Chua NH, and Carlier MF. Role of nucleotide exchange and hydrolysis in the function of profilin in action assembly. J. Biol. Chem. 1996; 271(21):12302-9. [PMID: 8647830]
- Ressad F, Didry D, Egile C, Pantaloni D, and Carlier MF. Control of actin filament length and turnover by actin depolymerizing factor (ADF/cofilin) in the presence of capping proteins and ARP2/3 complex. J. Biol. Chem. 1999; 274(30):20970-6. [PMID: 10409644]
- Yarmola EG, Dranishnikov DA, and Bubb MR. Effect of profilin on actin critical concentration: a theoretical analysis. Biophys. J. 2008; 95(12):5544-73. [PMID: 18835900]
- Kang F, Purich DL, and Southwick FS. Profilin promotes barbed-end actin filament assembly without lowering the critical concentration. J. Biol. Chem. 1999; 274(52):36963-72. [PMID: 10601251]
- Didry D, Carlier MF, and Pantaloni D. Synergy between actin depolymerizing factor/cofilin and profilin in increasing actin filament turnover. J. Biol. Chem. 1998; 273(40):25602-11. [PMID: 9748225]


