What are cell-matrix receptors?
Interaction between cell-matrix receptors and their respective ligands are often the initial step in the formation of a cell-matrix adhesion. Several types of cell-matrix receptors have been identified, each interacting with a specific type of ligand. Attachment of these various ECM based ligands or molecules to the exterior portion of the adhesion receptor causes their structural rearrangement [1]. This may be induced by a specific chemical property or change, (reviewed in [2]), a change in topography [3][4][5] (reviewed in [6][7][8]), or even the rigidity of the ECM components [9][10].
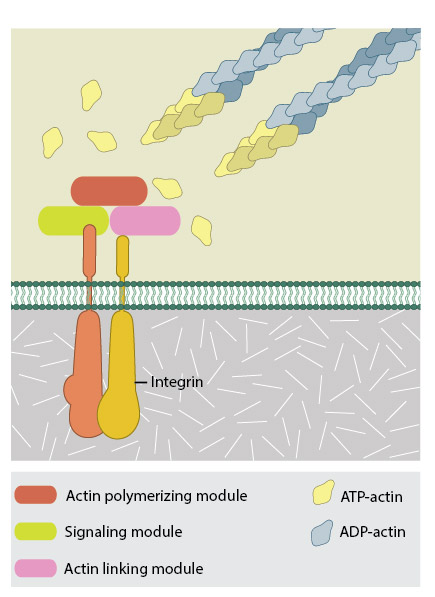
Integrins upon binding to ECM ligand gets activated and undergoes conformational changes. This leads to a series of events including i) recruitment of binding proteins to the site leading to actin linking, ii) activation of signaling molecules leading to integrin clustering and iii) actomyosin contractions leading to adhesion strengthening. All these result in iv) formation of new actin filaments by actin polymerization modules.Cell-matrix adhesion receptors are grouped according to the ligand that is bound. In some cases, adhesion receptors bind more than one type of ligand (e.g. the integrin family) and different receptor groups may cooperate with each other to bind their ligands (reviewed in [18583124, 19355970])
Various types of cell-matrix receptors exist. These include:
- Fibronectin receptors, which include the most common types of adhesion receptors, the integrin family and the syndecan family of transmembrane proteoglycans.
- Collagen receptors, which includes integrins, receptor tyrosine kinases (e.g. discoidin domain receptors [DDRs]), glycoproteins (e.g. GPVI) and immunoglobulin-like proteins aka IgCAMs (e.g. LAIR-1).
- Laminin receptors, which include integrins, the dystrophin glycoprotein complex (DGC), the 67 kDa laminin receptor (67LR), and two glycoproteins belonging to the immunoglobulin superfamily, Lutheran (Lu) and basal cell adhesion molecule (B-CAM).
- Hyaluronan receptors (aka hyaladherins) Proteins that bind to hyaluronan, a large polysaccharide, have immunoglobulin-like domains and include members of the CD44 family and CD168.
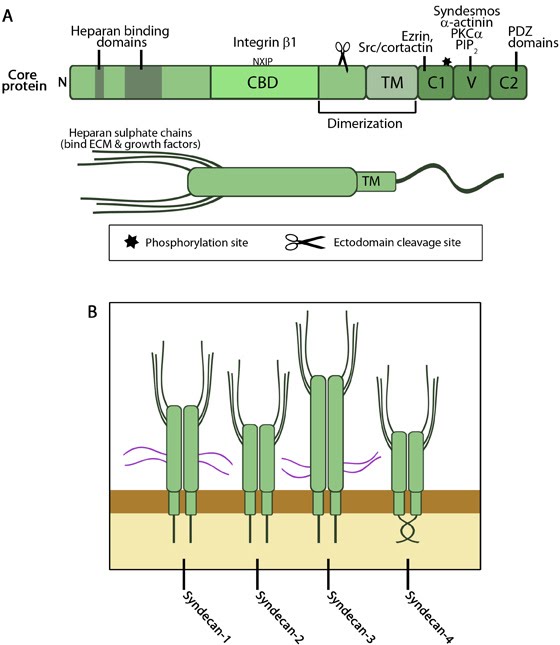
A. The extracellular domains of syndecan enable binding to heparan sulphate and integrin. The transmembrane domain drives dimerization, and the cytosolic domains promote binding to focal adhesion components. B. The four member of the syndecan family.
References
- Johnson CP, Tang H, Carag C, Speicher DW, and Discher DE. Forced unfolding of proteins within cells. Science 2007; 317(5838):663-6. [PMID: 17673662]
- Hersel U, Dahmen C, and Kessler H. RGD modified polymers: biomaterials for stimulated cell adhesion and beyond. Biomaterials 2003; 24(24):4385-415. [PMID: 12922151]
- Curtis A, and Wilkinson C. New depths in cell behaviour: reactions of cells to nanotopography. Biochem. Soc. Symp. 1999; 65:15-26. [PMID: 10320930]
- Dalby MJ, Riehle MO, Johnstone H, Affrossman S, and Curtis ASG. In vitro reaction of endothelial cells to polymer demixed nanotopography. Biomaterials 2002; 23(14):2945-54. [PMID: 12069336]
- Parker KK, Brock AL, Brangwynne C, Mannix RJ, Wang N, Ostuni E, Geisse NA, Adams JC, Whitesides GM, and Ingber DE. Directional control of lamellipodia extension by constraining cell shape and orienting cell tractional forces. FASEB J. 2002; 16(10):1195-204. [PMID: 12153987]
- Vogel V, and Sheetz M. Local force and geometry sensing regulate cell functions. Nat. Rev. Mol. Cell Biol. 2006; 7(4):265-75. [PMID: 16607289]
- Curtis A, and Riehle M. Tissue engineering: the biophysical background. Phys Med Biol 2001; 46(4):R47-65. [PMID: 11324976]
- Spatz JP, and Geiger B. Molecular engineering of cellular environments: cell adhesion to nano-digital surfaces. Methods Cell Biol. 2007; 83:89-111. [PMID: 17613306]
- Discher DE, Janmey P, and Wang Y. Tissue cells feel and respond to the stiffness of their substrate. Science 2005; 310(5751):1139-43. [PMID: 16293750]
- Engler AJ, Sen S, Sweeney HL, and Discher DE. Matrix elasticity directs stem cell lineage specification. Cell 2006; 126(4):677-89. [PMID: 16923388]
- Smith A, Sengupta K, Goennenwein S, Seifert U, and Sackmann E. Force-induced growth of adhesion domains is controlled by receptor mobility. Proc. Natl. Acad. Sci. U.S.A. 2008; 105(19):6906-11. [PMID: 18463289]
- Cavalcanti-Adam EA, Volberg T, Micoulet A, Kessler H, Geiger B, and Spatz JP. Cell spreading and focal adhesion dynamics are regulated by spacing of integrin ligands. Biophys. J. 2007; 92(8):2964-74. [PMID: 17277192]
- Lock JG, Wehrle-Haller B, and Strömblad S. Cell-matrix adhesion complexes: master control machinery of cell migration. Semin. Cancer Biol. 2007; 18(1):65-76. [PMID: 18023204]
- Gupton SL, and Waterman-Storer CM. Spatiotemporal feedback between actomyosin and focal-adhesion systems optimizes rapid cell migration. Cell 2006; 125(7):1361-74. [PMID: 16814721]
- Keren K, Pincus Z, Allen GM, Barnhart EL, Marriott G, Mogilner A, and Theriot JA. Mechanism of shape determination in motile cells. Nature 2008; 453(7194):475-80. [PMID: 18497816]
- Geiger B, Spatz JP, and Bershadsky AD. Environmental sensing through focal adhesions. Nat. Rev. Mol. Cell Biol. 2009; 10(1):21-33. [PMID: 19197329]


